This tutorial focus on the implementation of the image segmentation architecture called UNET in the PyTorch framework. It’s a simple encoder-decoder architecture developed by Olaf Ronneberger et al. for Biomedical Image Segmentation in 2015 at the University of Freiburg, Germany.
What is Image Segmentation?
An image consists of multiple objects inside it, such as people, cars, animals, or any other object. To classify the image, we use image classification, where the task is to predict the label or class of the input image. Now imagine, we need to find the exact location of the object, i.e, which pixel belongs to the which object. In this case, we want a pixel-level classification, i.e, we want to segment the image.
So, image segmentation is the process where a network takes an image as input and outputs a pixel-wise mask. This helps in a better understanding of the scene in the image at the pixel level. Image segmentation widely used in medical imaging, autonomous vehicle, satellite imaging, and many more.
UNET – Network Architecture
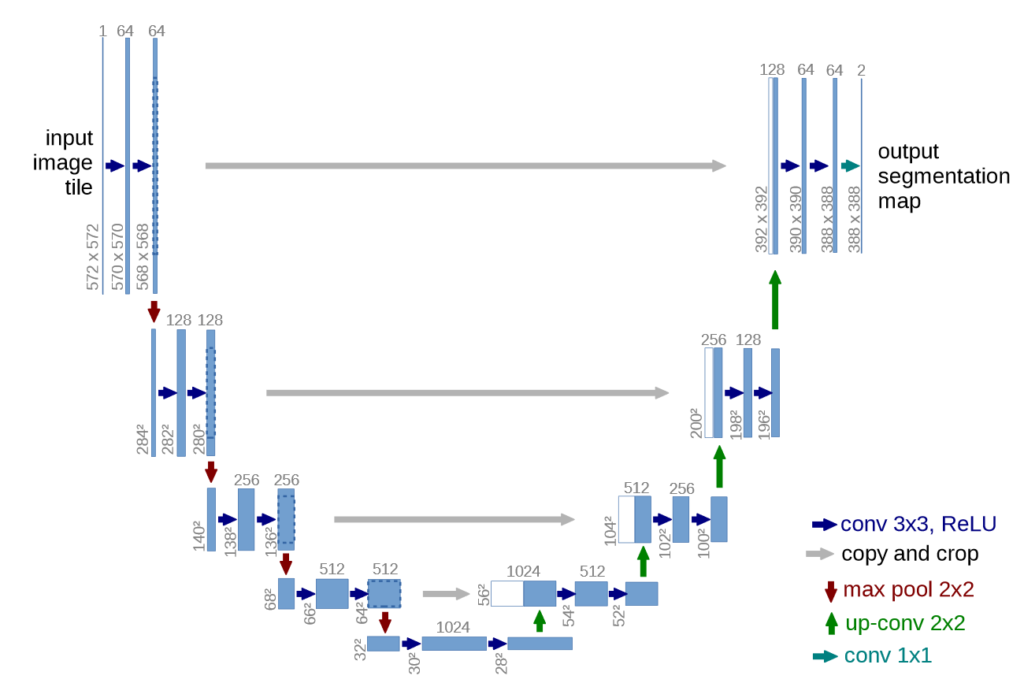
UNET is a U-shaped encoder-decoder network architecture, which consists of four encoder blocks and four decoder blocks that are connected via a bridge. The encoder network (contracting path) half the spatial dimensions and double the number of filters (feature channels) at each encoder block. Likewise, the decoder network doubles the spatial dimensions and half the number of feature channels.
READ MORE: What is UNET?
Import Torch
import torch
import torch.nn as nnConvolutional Block
The entire UNET architecture uses two 3×3 convolutional layers, each followed by a ReLU activation.
Here we creates a simple class named conv_block.
class conv_block(nn.Module):
def __init__(self, in_c, out_c):
super().__init__()
self.conv1 = nn.Conv2d(in_c, out_c, kernel_size=3, padding=1)
self.bn1 = nn.BatchNorm2d(out_c)
self.conv2 = nn.Conv2d(out_c, out_c, kernel_size=3, padding=1)
self.bn2 = nn.BatchNorm2d(out_c)
self.relu = nn.ReLU()
def forward(self, inputs):
x = self.conv1(inputs)
x = self.bn1(x)
x = self.relu(x)
x = self.conv2(x)
x = self.bn2(x)
x = self.relu(x)
return xThe original UNET does not use batch normalization in between the convolution layer and ReLU. Here we insert batch normalization in between them. It helps the network to reduces internal covariance shift and makes the network more stable while training.
Encoder Block
From the original paper
The contracting path follows the typical architecture of a convolutional network. It consists of the repeated application of two 3×3 convolutions (unpadded convolutions), each followed by a rectified linear unit (ReLU) and a 2×2 max pooling operation with stride 2 for downsampling. At each downsampling step we double the number of feature channels.
U-Net: Convolutional Networks for Biomedical Image Segmentation – Olaf Ronneberger et. al
class encoder_block(nn.Module):
def __init__(self, in_c, out_c):
super().__init__()
self.conv = conv_block(in_c, out_c)
self.pool = nn.MaxPool2d((2, 2))
def forward(self, inputs):
x = self.conv(inputs)
p = self.pool(x)
return x, pIn the encoder_block, we have used padding to make sure that the shape (height and width) of the output feature maps remain same as the input feature maps.
The encoder_block consists of an conv_block followed by a 2×2 max pooling. The number of filters are doubled and the height and width is reduced half after every block.
The encoder_block return two output:
- x: It is the output of the conv_block and acts as the input of the pooling layer and as the skip connection feature map for the decoder block.
- p: It is the output of the pooling layer.
Decoder Block
From the original paper
Every step in the expansive path consists of an upsampling of the feature map followed by a 2×2 convolution (“up-convolution”) that halves the number of feature channels, a concatenation with the correspondingly cropped feature map from the contracting path, and two 3×3 convolutions, each followed by a ReLU
U-Net: Convolutional Networks for Biomedical Image Segmentation – Olaf Ronneberger et. al
In the above lines, the up-convolution is referred to the transpose convolution.
class decoder_block(nn.Module):
def __init__(self, in_c, out_c):
super().__init__()
self.up = nn.ConvTranspose2d(in_c, out_c, kernel_size=2, stride=2, padding=0)
self.conv = conv_block(out_c+out_c, out_c)
def forward(self, inputs, skip):
x = self.up(inputs)
x = torch.cat([x, skip], axis=1)
x = self.conv(x)
return xUNET Architecture
class build_unet(nn.Module):
def __init__(self):
super().__init__()
""" Encoder """
self.e1 = encoder_block(3, 64)
self.e2 = encoder_block(64, 128)
self.e3 = encoder_block(128, 256)
self.e4 = encoder_block(256, 512)
""" Bottleneck """
self.b = conv_block(512, 1024)
""" Decoder """
self.d1 = decoder_block(1024, 512)
self.d2 = decoder_block(512, 256)
self.d3 = decoder_block(256, 128)
self.d4 = decoder_block(128, 64)
""" Classifier """
self.outputs = nn.Conv2d(64, 1, kernel_size=1, padding=0)
def forward(self, inputs):
""" Encoder """
s1, p1 = self.e1(inputs)
s2, p2 = self.e2(p1)
s3, p3 = self.e3(p2)
s4, p4 = self.e4(p3)
""" Bottleneck """
b = self.b(p4)
""" Decoder """
d1 = self.d1(b, s4)
d2 = self.d2(d1, s3)
d3 = self.d3(d2, s2)
d4 = self.d4(d3, s1)
""" Classifier """
outputs = self.outputs(d4)
return outputsThe code above the complete implementation of the UNET architecture in the PyTorch framework.