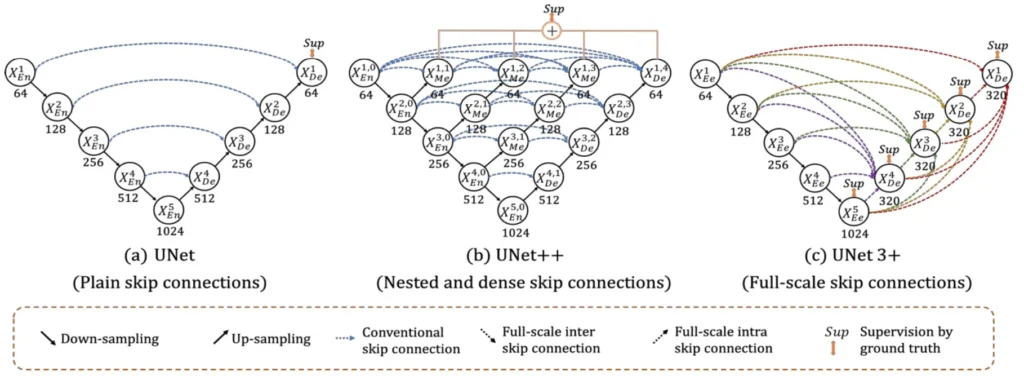
In medical image analysis, accurately identifying and outlining organs is vital for clinical applications such as diagnosis and treatment planning. The UNet architecture, a widely favoured choice for these tasks, has seen enhancements through UNet++, which introduced nested and dense skip connections to improve performance. Taking this evolution further, the introduction of UNet 3+ marks a significant advancement over UNet++, presenting an innovative approach to overcome current limitations and elevate segmentation accuracy to unprecedented levels.
Research Paper: UNET 3+: A FULL-SCALE CONNECTED UNET FOR MEDICAL IMAGE SEGMENTATION
What is UNet 3+?
UNet 3+ is a U-shape encoder-decoder architecture built upon the foundation of its predecessors, i.e., UNet and UNet++. It aims to capture both fine-grained details and coarse-grained semantics from full scales. The paper highlights the re-design of inter and intra-connections between the encoder and the decoder, providing a more comprehensive understanding of organ structures. Additionally, a hybrid loss function contributes to accurate segmentation, especially for organs appearing at varying scales, while reducing network parameters to improve computational efficiency.
Issues with Existing Methods
Existing segmentation methods often face challenges in reducing false positives, especially in non-organ images. Traditional approaches employ attention mechanisms or predefined refinement methods like Conditional Random Fields (CRF) at inference. UNet 3+ takes a different route by introducing a classification task to predict the presence of organs in the input image, offering valuable guidance to the segmentation task.
Read More:
Proposed Method: UNet 3+
UNet 3+ combines the multi-scale features by re-designing skip connections as well as utilizing full-scale deep supervision, which provides fewer parameters but yields a more accurate position-aware and boundary-enhanced segmentation map.
It consists of the following three components:
- Full-scale skip connections
- Full-scale deep supervision
- Classification-guided Module (CGM)
Full-scale Skip Connections
UNet 3+ addresses the shortcomings of UNet and UNet++ by incorporating both smaller- and same-scale feature maps from the encoder and larger-scale feature maps from the decoder at each decoder layer. This innovative approach captures fine-grained details and coarse-grained semantics in full scales, enhancing the model’s position-awareness and boundary definition. Importantly, UNet 3+ achieves this with fewer parameters, ensuring computational efficiency.
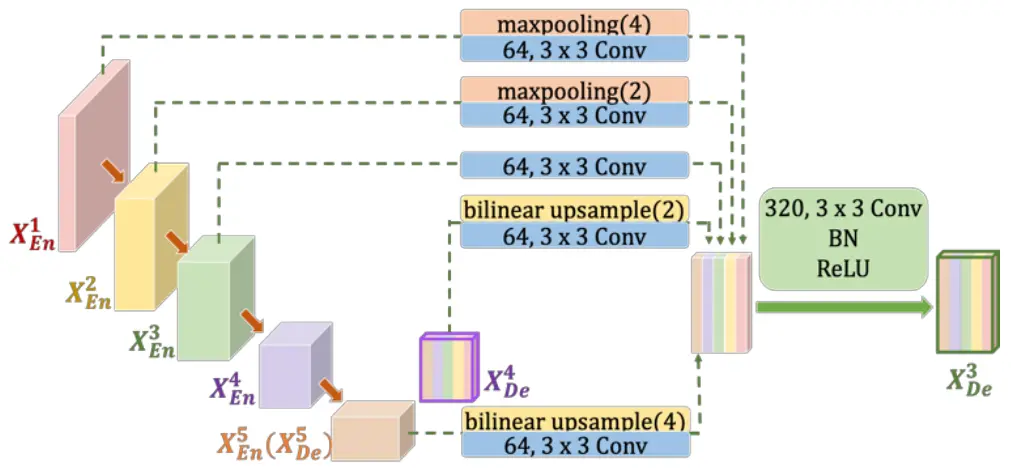
Full-Scale Deep Supervision
The adoption of full-scale deep supervision in UNet 3+ allows for better segmentation performance. The approach involves multiple supervision signals from different scales, contributing to the accurate delineation of organ boundaries.

Classification-guided Module (CGM)
To tackle false positives, particularly in non-organ images, UNet 3+ introduces a classification-guided module. This involves an additional classification task to predict organ presence in the input image. The classification result guides each segmentation side output, effectively addressing over-segmentation issues by providing corrective guidance.
Loss Function
The hybrid loss function employed by UNet 3+ includes a multi-scale structural similarity index (MS-SSIM) loss to monitor fuzzy boundaries. The total loss comprises Focal Loss, MS-SSIM Loss, and Intersection over Union (IoU) Loss. This comprehensive loss function operates at pixel, patch, and map levels, capturing both large-scale and fine structures with clear boundaries.
Total Loss = Focal Loss + MS-SSIM Loss + IoU Loss
Dataset and Implementation
The evaluation of UNet 3+ utilizes liver and spleen datasets obtained from the ISBI LiTS 2017 Challenge and an ethically approved hospital dataset. Input images are resized to 320 x 320 pixels. Stochastic Gradient Descent (SGD) is employed as the optimizer, with the dice coefficient serving as the evaluation metric.
Results
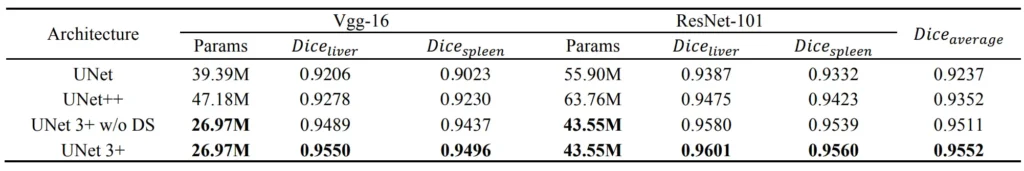

Conclusion
In conclusion, UNet 3+ presents a promising advancement in medical image segmentation, combining innovative architectural modifications, a hybrid loss function, and a classification-guided module. The results suggest improved accuracy, reduced false positives, and enhanced computational efficiency, making UNet 3+ a notable contribution to the field of medical image analysis.